Specialized iNANO Lecture by Professor Barbara Sacca,Center for Medical Biotechnology, University of Duisburg-Essen, Germany
Multivalent ligand-protein interactions in DNA-confined spaces
Info about event
Time
Location
1510-213 (AUD VI, Dept. of Chemistry)
Organizer
S. Gambietz, P. Stegemann, B. Möllers, M. Ehrmann and B. Saccà*, Center for Medical Biotechnology, University of Duisburg-Essen, Germany
*corresponding author: barbara.sacca@uni-due.de
Multivalent binding in nature occurs when multiple, tethered (and often identical) ligands target the surface of a receptor with a binding strength that is more than the sum of the individual components.[1] Spatial confinement instead can stabilize and accelerate folding of encapsulated proteins.[2] In this study, we merge these principles in programmable DNA-origami nanocompartments whose inner surfaces present precisely arranged, identical non-covalent ligands. By constructing oligomeric chambers of varied size and geometry and systematically tuning valency, ligand spacing and length, we create model systems in which stoichiometry and nanoscale architecture are defined a priori. Within these confined volumes we observe selective protein encapsulation with up to 1000-fold increase in apparent affinity, strongly dependent on both the number of ligands and their geometric arrangement. Binding is further governed by electrostatic interactions between
the protein surface and the negatively charged DNA shell, revealing a sweet-spot regime that maximizes encapsulation efficiency. We show that the protein’s effective surface charge can be modulated via fluorophore labeling at reactive residues, while sodium chloride addition mitigates nonspecific interactions by ionic screening. Together, these results decouple and quantify the contributions of multivalency, geometry, and
electrostatics in ligand-protein recognition, thereby contributing not only to a deeper understanding of multivalency in confined spaces, but also to the development of powerful synthetic tools for addressing specific biological questions with molecular detail.
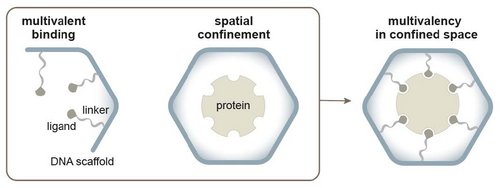
Figure 1. Multivalent binding relies on many identical ligands anchored to the same scaffold through linkers. Spatial confinement reduces the entropy of the system by limiting the probability
of the protein to assume unfolded conformations. Merging both phenomena in one single system may favor the formation of the ligand/protein complex and its thermodynamic stability.
REFERENCES:
[1] aM. Mammen, S. K. Choi, G. M. Whitesides, Angew Chem Int Ed Engl 1998, 37, 2754–2794; bP.
I. Kitov, D. R. Bundle, J Am Chem Soc 2003, 125, 16271–16284.
[2] H. X. Zhou, K. A. Dill, Biochemistry-Us 2001, 40, 11289–11293.