AU researchers contribute to understanding corona proteins on nanoparticles
The properties of nanoparticles are widely acknowledged and they are an important tool in pharmaceutical applications, among others. However, there is a need for deeper understanding of the protein layers accumulating on their surface, as these protein layers affect the functional role of the nanoparticles. AU researchers have developed a method for more efficient studies of the protein composition, which recently has been published in Nature Communications.
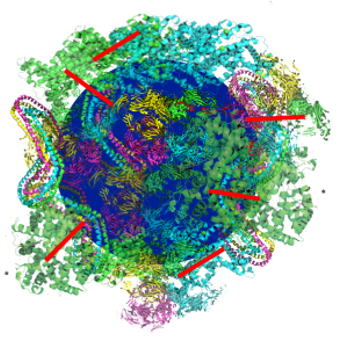
Nanoparticles (NPs) are promising agents for drug delivery and visualization of processes in live cells. When adding NPs to a biofluid (blood, urine, etc.) proteins present in the specific biofluid attach to the surface of the nanoparticle, and a layer of proteins, “corona proteins”, arise on the NPs surface. These corona proteins play a major role in the binding of NPs to cell surfaces and thus have a strong impact on the function of NPs.
The corona proteins are proteins that competitively adhere to the surface of nanoparticles to form a combined “Hard” (HC) and “Soft” corona (SC). HC proteins with a high binding affinity and low dissociation rate remain tightly bound to the surface, whereas SC proteins with a high dissociation rate are rapidly exchanged.
Missing links in the protein layer of nanoparticles
Several researchers have studied the protein corona formation and identification of the proteins forming the corona. The research has been aiming to understand how NPs interact with cell surfaces, nanoparticle toxicity, etc. However, there are still several missing links in connection to understanding the complex and dynamic process of proteins adhering to the NPs.
The reason for this is that the current understanding of the biological roll that nanoparticles may acquire in a given biological milieu is mostly inferred from the hard component of the protein corona (HC). The composition of soft corona (SC) proteins and their biological relevance has remained elusive due to the lack of analytical separation methods.
Weakly interacting corona proteins may have an important role
AU researchers report of a new approach to studying the corona proteins in Nature Communications with Dr. Hossein Mohammad-Beigi as first-author (member of Duncan Sutherland's research group).
Here, the researchers have identified a set of specific corona proteins with weak interactions at silica and polystyrene nanoparticles by using an experimental approach based on click chemistry. It appears that these SC proteins are present also in the HC, but are specifically enriched after the capture, suggesting that the main distinction between HC and SC is the differential binding strength of the same proteins.
Recent work has developed approaches aimed at quantifying and identifying SC and HC proteins, but they are not yet effective. Here, the new approach is faster, clearer, and provides the possibility to study the effect of the dynamic nature of the weakly interacting SC proteins on cell association along with the long-lived protein corona forming around nanoparticles in complex media.
Interestingly, the SC proteins are revealed as modulators of nanoparticle-cell association mainly through their dynamic nature, as the results show that the NP surface, though covered by corona protein, still can interact directly with a cell surface. This suggests that the properties of the bare particle surface can still directly influence cell association even after a fully developed protein corona is formed.
The researchers, therefore, highlight that weak interactions of proteins at nanoparticles should be considered when evaluating nano-bio interfaces.
Read more about the results in Nature Communications:
The research was carried out by researchers from Interdisciplinary Nanoscience Center (iNANO), the Department of Molecular Biology and Genetics, and Department of Biomedicine at Aarhus University. The work was financially supported by Independent Research Fund Denmark, Danish National Research Foundation (Centre for Cellular Signal Patterns, CellPat), and the Lundbeck Foundation.
For further information, please contact
Professor Duncan Sutherland
Interdisciplinary Nanoscience Center
Aarhus University
Email: duncan@inano.au.dk